The B.1.617 FAMILY OF COVID VARIANTS
The ongoing surge in India is a fertile breeding ground for new SARS-CoV-2 variants
THE B.1.617 FAMILY OF COVID VARIANTS
15 May 2021/Saturday/1230am CDT
“SARS-CoV-2, like other RNA viruses, is constantly generating mutations as it replicates inside infected people. The reason that SARS-CoV-2 has been accumulating many more mutations during the past several months is probably related to increasing spread and transmission around the world. The more viral replication occurring in the human population, the more random mutations are generated, and the greater the chance that one of these mutations will improve infection or transmission, resulting in a new viral variant that outcompetes earlier ones."
-Scott C. Weaver, Professor and Chair, Department of Microbiology and Immunology, Director, Institute for Human Infections and Immunity, Scientific Director, Galveston National Laboratory, University of Texas Medical Branch
High resolution version of today's pandemic graphic:
https://drive.google.com/file/d/1Unf7n6IodZ2Ue2AI67hVFFmoxbfIV7vp/view?usp=sharing
BACKGROUND
The pandemic situation in India has captured the world's attention for its rapid escalation to the point of overwhelming not just the medical infrastructure of the country, but even it's post-mortem infrastructure, with sobering images of numerous fires from outdoor temporary crematoriums to handle the cremation of the victims of COVID.
About a month ago, I wrote about how COVID case surges are fertile grounds for the emergence of COVID variants and showed how the variants we knew about then could be tied back to past case surges in the countries they emerged based on the genomic surveillance data (links to that past posting at the end of this post). At the time of my post, B.1.617 that emerged in India was not yet on the radar screen. I had seen reports in the Indian press about a new "double mutant".
“I think most people studying viral evolution agree that the appearance of these variants now mostly reflects the very large number of cumulative infections that have occurred worldwide since the start of the pandemic. On average, one mutation may occur every time or every other time the virus infects a new person. Each mutation is kind of like pulling a slot machine — the chance of hitting the jackpot on any individual pull is small, but you pull millions of handles simultaneously the chances are dramatically increased. Viruses that ‘hit the jackpot’ by accumulating a set of mutations that makes them more transmissible will then increase in the population due to natural selection."
-Thomas Friedrich, PhD , Professor, Department of Pathobiological Sciences, University of Wisconsin School of Veterinary Medicine
Each new person infected by the virus is an opportunity to mutate. Should that mutation offer some selective advantage to the virus, you will likely have the emergence of a new variant as it will have a fitness advantage over other variants and preferentially spread.
Six days after my posting on how the emergence of new variants could be linked to past case surges, a new variant of interest appeared that was first detected in India- B.1.617. This was the "double mutant" being reported in the Indian press starting at the end of March. (I have links to the post I wrote about B.1.617 at the end of this post).
Watching the data on the exponential growth of the cases and deaths in India from COVID, it was only a matter of time before more variants emerged out of the pandemic environment of thousands and thousands of new infections each day.
What I did not expect was that B.1.617 has split into three distinct variants. Well, not that unexpected- after all, we have precedent with B.1.526 that first emerged likely in New York that has split into sub-lineages, B.1.427 and B.1.429 that first emerged in California split from a common parent variant as did P.1 and P.2 in Brazil.
Two of the sub-variants, B.1.617.1 and B.1.617.3, are considered variants of interest and B.1.617.2 is considered a variant of concern given how rapidly it has become the dominant variant in COVID infections in India in the last several weeks.
I am in the process of updating (again) my COVID Variants Table. Stay tuned for that update.
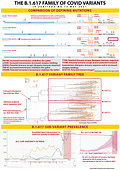
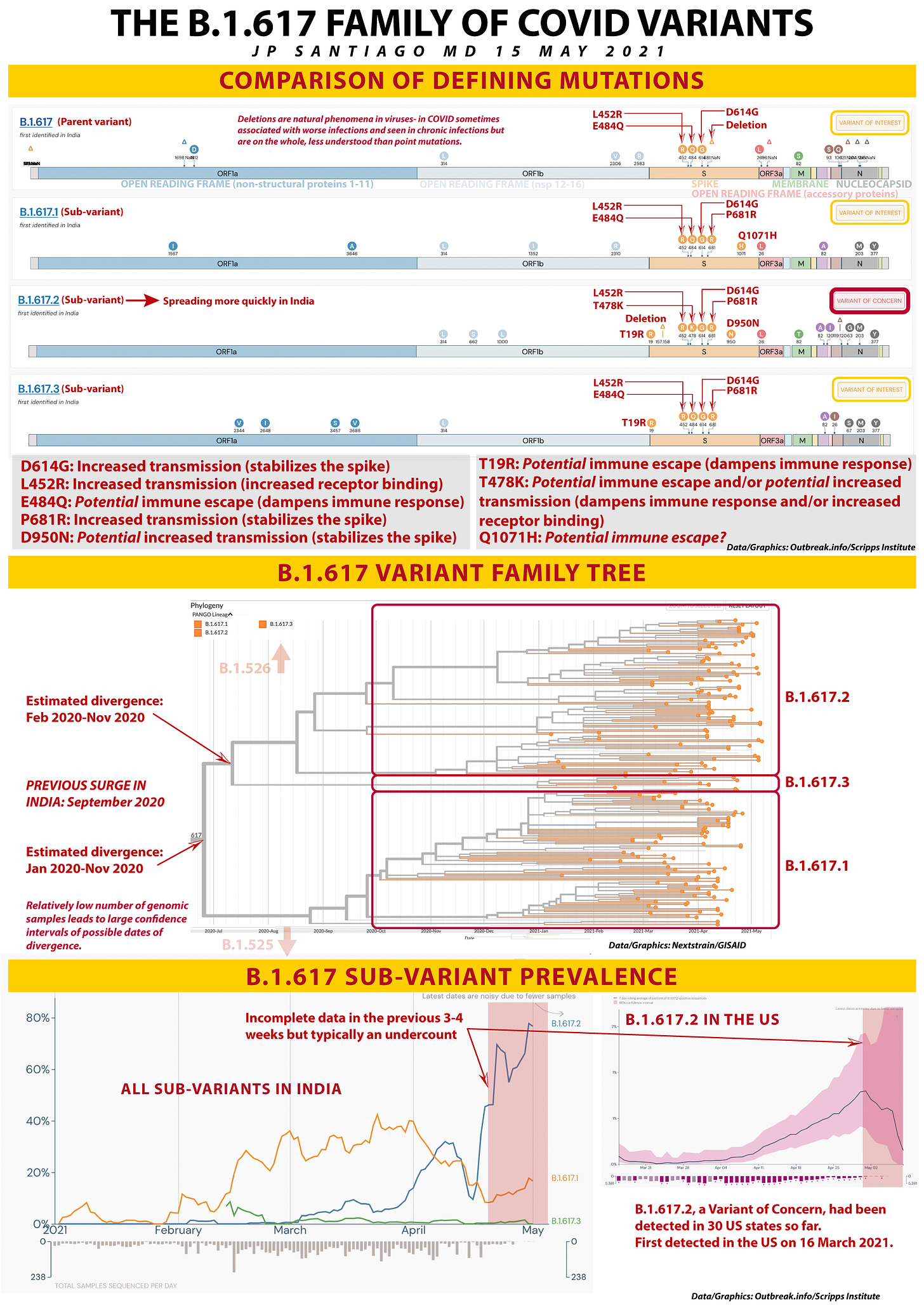
COMPARISON OF DEFINING MUTATIONS
The top part of today's pandemic graphic has four genomic diagrams representing B.1.617, the parent variant that I posted about on 18 April, and the three sub-variants, B.1.617.1, 617.2, and 617.3. Note that on the far right I have highlighted which ones are "Variants of Interest" and which one is the "Variant of Concern".
BUT FIRST, LET'S TALK ABOUT THE COVID GENOME
I have posted before that the SARS-CoV-2 (COVID) genome is about 30,000 nucleotides long. For comparison, the human genome is about 3 billion nucleotides long. We're a more complex organism, so we have tons more genes.
There are three kinds of genes in the viral genome- structural proteins, non-structural proteins, and accessory proteins.
One of the structural proteins we're all very aware of- the "S gene" which encodes for the spike. There is also the "M gene" for membrane proteins, "N gene" for the nucleocapsid proteins (these protect the viral genome), and the "E gene" for envelope proteins. Membrane and Envelope proteins form the spherical shell of the virus, the Spike proteins is what the virus uses to bind with the host receptor (ACE2 for COVID) and start the infection process.
On the genome diagrams, I have noted where the Spike, Membrane and Nucleocapsid genes are located. The Envelope gene is a small section on the right side.
Non-structural proteins are encodes in the ORF genes which stand for "Open Reading Frame". Non-structural proteins are used primarily to facilitate the viral replication process as the virus hijacks our cells to make millions of copies of itself to infect more cells in our bodies and other people around us. ORF1a and ORF1b encode for the non-structural proteins (or "nsp") on the left side of the viral genome. ORF1a encodes for eleven nsp's, ORF1b encodes for five nsp's.
Accessory proteins are the third kind of viral protein. They are encoded on short ORF genes on the right side like ORF3a. Accessory proteins are the least understood proteins but we know that some of them are involved with trying to inhibit or modulate our immune response to infection.
There are mutations that occur in the ORF genes and the structural protein genes, but they don't drive as much of the clinical and epidemiological behavior of the virus as much as mutations in the Spike gene. As I have posted previously, mutations in the Spike gene change the behavior of certain parts of the Spike- either increased binding (which means it's more contagious), increased stability (also makes it more contagious), or immune escape (it dampens our immune system's response to infection).
MUTATIONS OF NOTE
I have taken genome diagrams from Outbreak.info/Scripps Institute and annotated them with the key spike mutations.
The small triangles mark where deletion mutations have occurred. I have not really touched on deletions in my past posts. Deletions are where a short segment (sometimes as little as a single nucleotide) of the genetic code is removed. So instead of a nucleotide swap, it just gets deleted all together.
Deletions are a natural phenomena in all viruses, but in COVID it appears that deletions are likely associated with worse infections and are also associated with chronic infections. They are less understood than point mutations like L452R or E484Q.
So each variant genome diagram marks out mutations of note, most of what we are really interested in are spike mutations because those have the greatest impact on how the virus behaves in our patients and in populations.
There are several spike mutations I have noted:
D614G: This one emerged last spring and by the summer was present in the majority of viral genome samples worldwide. My past post on the molecular biology of the COVID spike discussed why D614G made COVID more contagious (links below at the bottom of this post).
L452R: This mutation in the receptor binding domain is also in B.1.427 and B.1.429 that emerged in California last year and drove their fourth quarter of 2020 surge. It makes COVID about 20% more contagious and this mutation is also present in B.1.526.1 that emerged in New York this past fall.
E484Q: This is in the same location as the E484K mutation I wrote about several days ago. It's in a location called an epitope which is a target for neutralizing antibodies either via vaccination or monoclonal antibody therapy. While it appears at the molecular level that E484Q is not as serious as E484K, it still represents a change on a vital target of our therapies against the virus.
P681R: Like D614G, this mutation more than likely serves to stabilize the spike in a conformation that makes it more likely and more efficient at infecting our cells.
D950N: I am still doing more reading on this mutation but it appears likely stabilize the spike in a more infectious configuration.
T19R: This one I just learned about yesterday. This mutation is at one tip of the spike not part of the receptor binding domain but is likely part of an important epitope. As such, it holds the potential for dampening our immune response.
T478K: This another interesting mutation- its location is not near the receptor binding domain which leads us to believe it may be part of an epitope. There was a variant that emerged in Mexico with T478K that rapidly became the majority of that country's cases this spring and it had T478K which suggests it may also impact receptor binding, making it more contagious. More work is ongoing on this mutation.
Q1071H: This mutation I am not certain about but it's location on the Spike suggests it may be part of an epitope.
B.1.617.2 has all these mutations except Q1071H. We know from the epidemiological data B.1.617.2 is spreading faster through India than its 617 siblings.
THE B.1.618 VARIANT FAMILY TREE
I pulled this phylogenetic tree off Nextstrain with an Asia-focused dataset in the GISAID genome database. I've marked out where the three sub-variants are located. Interestingly B.1.617.3 was the first one identified. Keep in mind that we only have less than 15,000 genomic samples from India and the fewer samples you have, the more provisional your phylogenetic tree is going to be.
As a result, the dates of divergence (when they emerged as a distinct variant) have large confidence intervals- nearly a year long! The more samples we get into the database from India, the smaller the confidence intervals and the more accurate the date of emergence. For reference, prior to the current COVID surge in India, the last surge there was in September 2020. If I were to suggest a potential date of emergence, that's one starting point.
The lightly shaded notations note the nearest "relatives" on the bigger phylogenetic tree are located. If we had the full family tree, B.1.526 (first emerged in New York) would be just above this group and B.1.525 (first emerged in Nigeria) would be below. So those variants of interest are more closely related to the B.1.617 variant family than say P.1 or B.1.351.
B.1.617 SUB-VARIANT PREVALENCE
I pulled the data for the two graphs here from Outbreak.info. The first one on the left is the sub-variant prevalence in India for the three B.1.617 siblings. Keep in mind that because genomic sequencing is a highly specialized process, it takes time to get that data, so any genomic surveillance data in the previous 3-4 weeks will still be incomplete. If anything, there is usually an undercount in the previous month.
You can clearly see that B.1.617.2 is rapidly increasing and dominant in India.
The graph on the right is B.1.617.2 prevalence in the United States. Still small numbers, but it has been detected already in 30 states and was first detected in the United States on 16 March 2021. At that time, it was not on everyone's radar screen as a variant of concern, the parent variant wasn't a variant of interest and it was only starting to be mentioned in the Indian press as the cases in the largest of the Indian provinces, Maharashtra, began skyrocketing.
TAKE HOME MESSAGES
1/ COVID surges continue to be fertile ground for the emergence of new variants. The pandemic environment in India with thousands and thousands of new infections each day has led to B.1.617 splitting into three genetically distinct sub-variants designated B.1.617.1, B.1.617.2, and B.1.617.3.
2/ Defining point mutations on the spike for B.1.617.2 make it a "Variant of Concern" as it has spread rapidly in India, outpacing the other 617 sub-variants. Some of the point mutations are known to increase how easily the virus is spread- D614G, L452R, while other point mutations have the possibility of immune escape or spike stabilization.
3/ B.1.617.2 has already been detected in 30 US states. On 28 April, BioNTech (Pfizer's partner in their mRNA vaccine) reported that their vaccine should be protective against the variants with the E484Q mutation. This was announced that day by BioNTech's CEO on German radio. That data is being generated as we speak.
PARTING THOUGHTS
My parting thoughts on my posting about B.1.617 last month bear repeating here:
It behooves us to assist all nations with their vaccination and pandemic mitigation efforts. COVID does not respect national borders and time and time again in this pandemic, air travel has been the means by which new variants have spread around the world. This is not to say that the airline industry is to blame- origin and destination countries on trunk airline routes need to have robust surveillance and mitigation infrastructure in place to slow the spread of new variants.
These surges that are going on around the world are creating favorable environments for new variants to emerge that I'm almost certain we'll find through the rest of this year as sequencing and surveillance efforts ramp up worldwide.
It's not enough to develop new vaccines. We also need ways of delivering and administering effective vaccines throughout the world. As it stands right now, mRNA vaccines for all their effectiveness are not well suited for distribution and administration in many nations. I am concerned that there will be this two-tier system of vaccination in the world- mRNA vaccines for the industrialized nations and vector vaccines for the developing nations. It smacks of colonialism in medical form when a vaccine we won't use in the United States is all that's available for developing nations.
I also had this to say in my post about the E484K mutation and it bears repeating particularly in light of the recent controversial changes in CDC guidance for those vaccinated vs. those not vaccinated:
Our best chance at limiting the emergence of new mutations and new variants is to limit the opportunities for mutation with each new infected individual. We have to absolutely control spread and limit new infections. Our best defense is still vaccination. Non-pharmacologic interventions at the public level like mask mandates, social distancing measures and capacity restrictions as well as improved ventilation of indoor spaces buy us crucial time to get as many people vaccinated as possible.
Don't give away your shot. There are millions of people in a lot of places worldwide that dearly wish to be in your position to have such ready access to COVID vaccinations.
RELEVANT PAST POSTS (FACEBOOK LINKS)
THE MOLECULAR BIOLOGY OF THE E484K MUTATION (10 May 2021): https://www.facebook.com/jp.j.santiago/posts/10218832046529173
IMMUNOLOGY 101: THE IMMUNE SYSTEM- REPOST/UPDATE (05 May 2021): https://www.facebook.com/jp.j.santiago/posts/10218801066954703
QUICK UPDATE: AMERICAN INDIANS/ALASKA NATIVES LEAD THE WAY ON VACCINATIONS (29 April 2021): https://www.facebook.com/jp.j.santiago/posts/10218763919866049
QUICK UPDATE: BY THE NUMBERS (27 April 2021): https://www.facebook.com/jp.j.santiago/posts/10218750780817581
COVID VARIANTS OVERVIEW AND UPDATE (22 April 2021): https://www.facebook.com/jp.j.santiago/posts/10218722676514991
NEW COVID VARIANT OF INTEREST: B.1.617 (18 April 2021): https://www.facebook.com/jp.j.santiago/posts/10218699298170547
CASE SURGES CAN RESULT IN COVID VARIANTS (12 April 2021): https://www.facebook.com/jp.j.santiago/posts/10218657527966318
THE OVERVIEW & STATUS OF COVID VARIANTS (9 April 2021): https://www.facebook.com/jp.j.santiago/posts/10218637120816152
MAYO CLINIC GRAND ROUNDS: COVID TESTING (12 March 2021): https://www.facebook.com/jp.j.santiago/posts/10218453381862793
QUICK UPDATE: THE COVID FAMILY TREE (01 February 2021): https://www.facebook.com/jp.j.santiago/posts/10218211054244754
COVID MUTANTS: VARIANTS OF CONCERN (31 January 2021): https://www.facebook.com/jp.j.santiago/posts/10218204381877949
COVID MOLECULAR BIOLOGY: THE SPIKE (6 January 2021): https://www.facebook.com/jp.j.santiago/posts/10218019683340601